Evolutionary genetics and genomics at the University of Montana
We are a collaborative team of researchers focused on addressing questions in the broader fields of evolutionary genetics and endosymbiosis. Most of our current projects integrate approaches from genomics, genetics, cell biology, and population biology to better understand interactions between endosymbiotic Wolbachia bacteria and their divergent insect hosts. We are specifically interested in explaining how/why Wolbachia became the most common endosymbionts in nature.
While our work mostly focuses on natural systems, it also contributes to improving human health. When placed into mosquitoes, the Wolbachia strains that our research explores can block viruses that cause human disease (particularly dengue and Zika). This approach is projected to protect 500 million individuals from disease by 2030. Very recently, we have also begun collaborative work focused on better understanding SARS-CoV-2 evolution. All of our work is only made possible due to the generous support of the National Institutes of Health and the National Science Foundation that have consistently funded our lab at the University of Montana since 2017.
While our work mostly focuses on natural systems, it also contributes to improving human health. When placed into mosquitoes, the Wolbachia strains that our research explores can block viruses that cause human disease (particularly dengue and Zika). This approach is projected to protect 500 million individuals from disease by 2030. Very recently, we have also begun collaborative work focused on better understanding SARS-CoV-2 evolution. All of our work is only made possible due to the generous support of the National Institutes of Health and the National Science Foundation that have consistently funded our lab at the University of Montana since 2017.
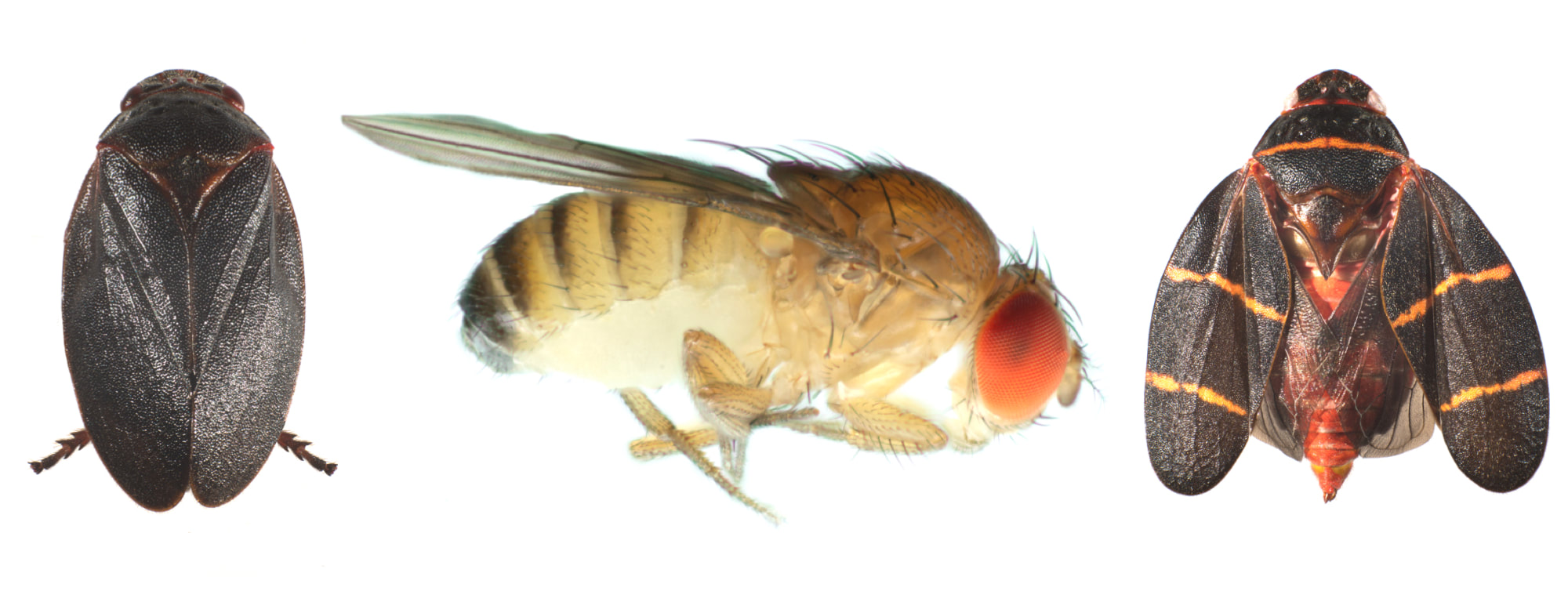
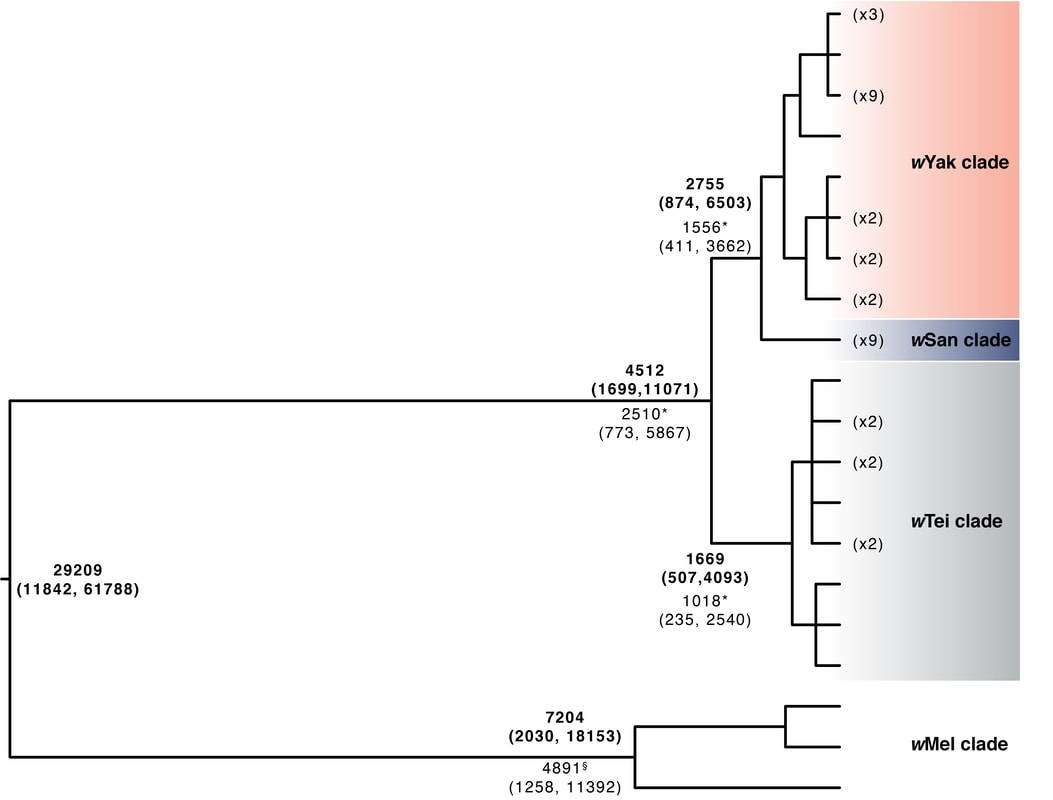
How do endosymbionts spread among host species?Wolbachia infect most insects, plus many other arthropods and some nematodes. We are interested in the modes and mechanisms of interspecific spread. To answer this question, we employ phylogenomic approaches that compare host and Wolbachia relationships and divergence times to test among three hypotheses: Wolbachia acquisition from common ancestors during speciation, from sister species due to hybridization, or from any number of divergent species via horizontal transfer. Surprising to us at least, Wolbachia seem to switch host on very short timescales via non-sexual transfer, with most hosts acquiring Wolbachia horizontally from distantly related taxa. We are interested in carrying these types of analyses out across even more divergent hosts to gain a better understanding of Wolbachia proliferation in nature.
|
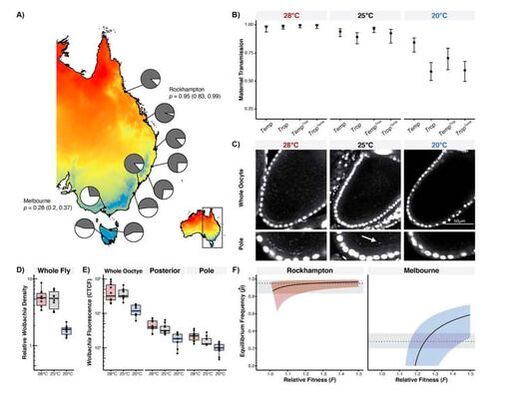
What are the causes and consequences of variable endosymbiont prevalence in host populations?While Wolbachia infect most insects, infections are often polymorphic in host populations. Wolbachia that cause intense cytoplasmic incompatibility (CI) tend to spread to high frequencies that are relatively stable, although not always. (CI reduces the egg hatch of uninfected females mated with Wolbachia-infected males.) In contrast, infections that do not cause CI or cause only weak CI occur at intermediate frequencies that often vary. Our research is determining how much and precisely why Wolbachia frequencies vary. Some of our most recent work has demonstrated how temperature effects on Wolbachia abundance in developing female oocytes influences the efficiency of Wolbachia transmission. Using mathematical models, we demonstrated that this type of transmission breakdown can explain variable Wolbachia prevalence in host populations, on continent-wide scales.
|
What underlies variation in how strongly endosymbionts influence host reproduction?
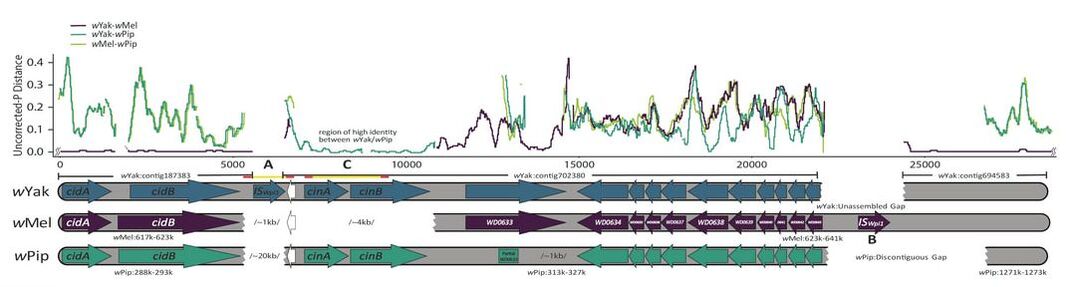
Much of our recent research is evaluating the evolutionary genetic basis of Wolbachia-induced CI. This work has included genomic analyses characterizing genetic variation associated with CI loci (cifs). For example, the figure below shows how sister Wolbachia may carry divergent cif copies that they can acquire horizontally. Our recent work has leverage these specific genomic results to design constructs that we express transgenically in yeast and fly—our efforts have revealed how single mutations can dramatically influence CI strength through effects on enzyme activity and efficiency. More broadly, we are characterizing the causes of CI-strength variation across divergent host systems. This work is revealing that while many factors may influence CI strength, the expression of a single gene (cifB) tends to explain most variation in CI.
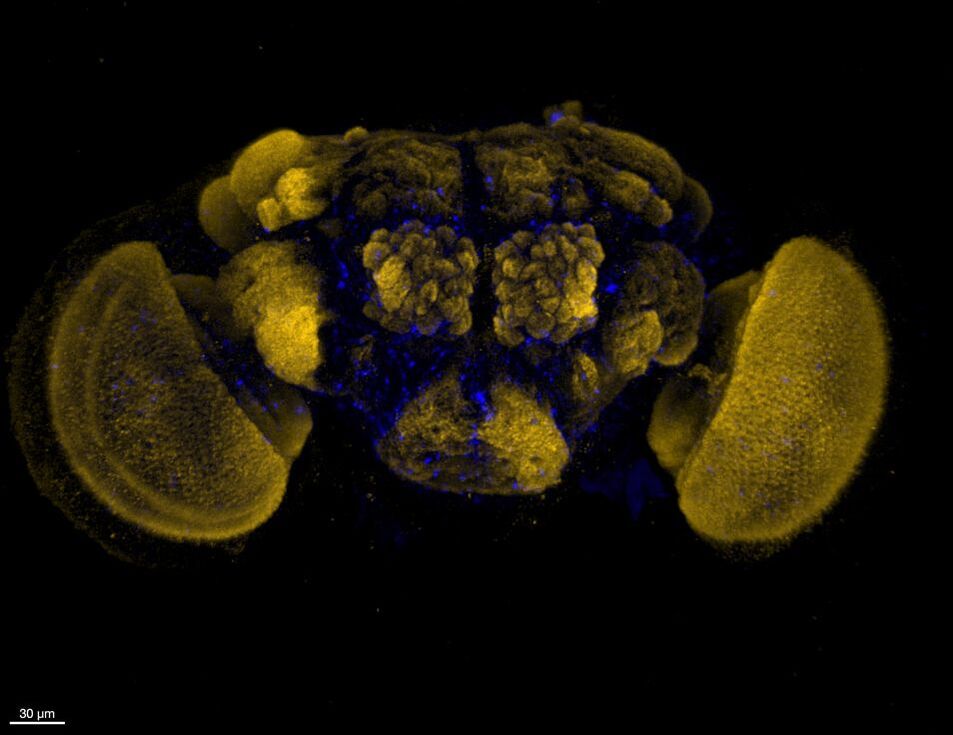
How much does endosymbiont abundance differ among host tissues, and what are the consequences for hosts?Many results from our lab and others have highlighted that the abundance and distribution of Wolbachia in host tissues can vary widely. We are interested in understanding how this variation influences both Wolbachia transmission and their effects on hosts. This work includes assessing variation in Wolbachia abundance and localization in developing female oocytes, in male testes, and in somatic tissues—we are particularly interested in identifying the genes that modify these traits. Our recent results have also demonstrated that Wolbachia infections covary with behavioral changes in hosts (e.g., thermoregulation and activity). Work in the lab is focused on connecting these behavioral phenotypes to Wolbachia presence and abundance in host tissues (preliminary brain image by John Statz).
|
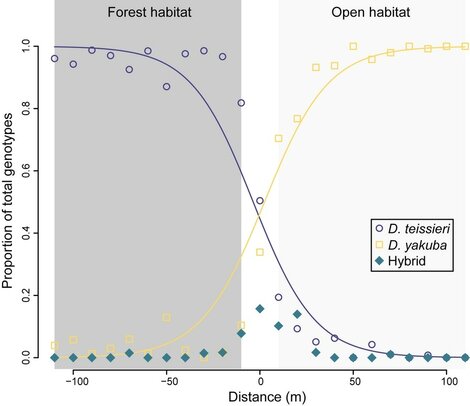
Local adaptation and reproductive isolation (Cooper's other life)I am also broadly interested in understanding local adaptation within, and the evolution of reproductive isolation between, host species. In the past this research has involved assessing the cell basis of temperature adaptation in natural and experimentally evolved Drosophila populations. More recently, I have collaborated with Daniel Matute on several projects, including identifying a D. yakuba-D. teissieri hybrid zone (and what maintains it) in west Africa. Because Wolbachia are maternally transmitted endosymbionts, hybridization and introgression can lead to between species Wolbachia spread. In this way, my work on local adaptation and reproductive isolation overlaps nicely with the lab's main work on Wolbachia. See my recent review in Evolution with Daniel if you are interested. Note that, most evidence suggests little contribution of Wolbachia to reproductive isolation, but there are interesting examples.
|
Current Funding
We are currently funded by a National Institutes of Health (NIH) R35 MIRA Award (2022-2027), a National Science Foundation CAREER Award (2022-2027), and by NIH funding in collaboration with Prof. Jeffrey Good and others to work on SARS-CoV-2 evolution in Montana and elsewhere (2022-23).